
Chapter 12 Add interactive tutorials
12.1 PART 1: Including a LearnR tutorial
12.1.1 Why do this?
LearnR tutorials are interactive R Markdown documents that allow you to incorporate code exercises as well as elements like videos and quizzes. There are many possibilities, and we point you to other resources to see various examples of these tutorials in action. The nice thing about this tool is that students can play around with code and preview various concepts outside of the RStudio IDE. One possible use case would be when you would like to introduce broader concepts in an interactive way before students begin focusing on the programming and coding aspects of the concepts within a regular .Rmd document.
12.1.2 Step 1: Create files and directories
As before, I assume you have at a minimum created the basic package structure from Part 1.
- Install the
learnrpackage - Run
use_tutorial("<name-of-tutorial-file>", "<title-you'd-like-the-user-to-see>", open = interactive())
remotes::install_github("rstudio/learnr")
use_tutorial("Lesson1", "My Title", open = interactive())This adds a tutorial folder to the inst directory.
12.1.3 Step 2: Customize your tutorial
- The
.Rmdfile inside thetutorialfolder is automatically opened. Edit it to customize your tutorial content.
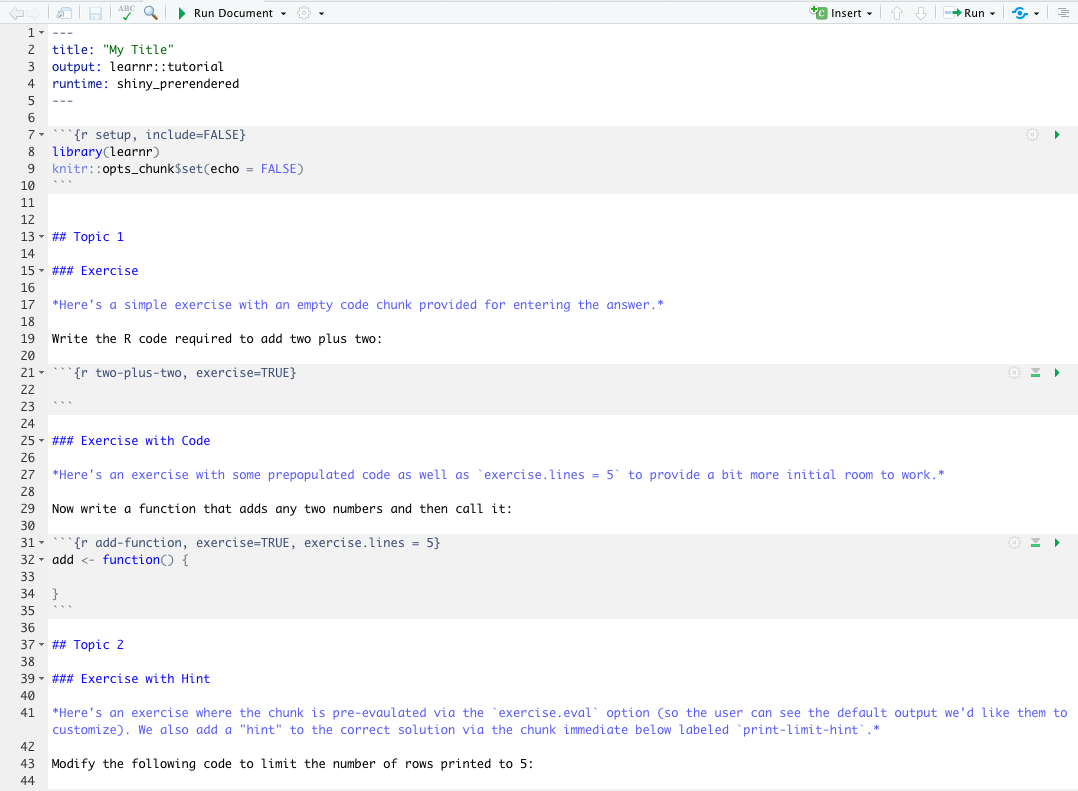
- Click Run Document next to the little green arrow in the toolbar of the
.Rmdfile.- This will generate a
.htmlfile in the same directory as the tutorial.Rmdfile and also run your tutorial locally.
- This will generate a
12.1.4 Step 3: Add additional subdirectory
At the time of writing, there is one additional folder that needs to be added, which use_tutorial did not create for us.
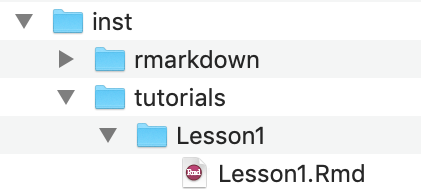
- Within the subdirectory
tutorials/, add a new folder with the same name as your.Rmdtutorial file. - Move your tutorial
.Rmdfile and the.htmlfor your tutorial into this new folder. - Build > Install and Restart
12.1.5 Step 4: Run your tutorial
- Run your tutorial with
run_tutorial("<tutorial-name>", "<package-name>").
learnr::run_tutorial("Lesson1", "testpackage")The above line of code is also how your users will be able to run your tutorial once they have installed your package.
If you get errors, double check that your structure for the tutorial folders and files matches that of those in the testpackage repository. Make sure you Build, Install, and Restart before trying again.
12.2 PART 5: FINAL STEPS
12.2.1 (Optionally) Document the package as a whole
We created documentation for the data set, but not for the package overall. To do so:
- Run
usethis::use_package_doc() - Edit the associated .R script that is generated in
R/with roxygen-style comments- See section 6.6 here and here for examples of what to include.
- Don’t forget to run
document()to update the documentation
#install.packages(c("devtools", "roxygen2"))
library(roxygen2)
library(devtools)
document()- Re-Build and install your package
- Check that package documentation works by typing
? <package-name>substituting your own package name.
12.2.2 Create a README.Rmd file
You can include additional information in a README.Rmd file for your package. At a minimum, you should include a line about how to install, using the guidelines from Part 1: Deliver.
- Run
use_readme_rmd()
> use_readme_rmd()
✔ Writing 'README.Rmd'
✔ Adding '^README\\.Rmd$' to '.Rbuildignore'
● Modify 'README.Rmd'
✔ Writing '.git/hooks/pre-commit'- Edit to meet your needs.
- Click
Knit, so that it updates the corresponding README.md file - Commit and push.
12.3 Troubleshooting
If you run into rough patches, here are some common pitfalls that might be resulting in errors:
- Do you have most up-to-date version of your packages? Try installing the development version of the package from github. Here’s an example of what that looks like for the
usethispackage.
# install.packages("devtools")
devtools::install_github("r-lib/usethis")- Are you running the most current version of R and/or RStudio?
- Did you install R via homebrew? –> Uninstall and install from CRAN!
12.4 Miscellaneous / Nice to Know
- If you didn’t start with a repo, then you should use
usethis::use_github() - Licensing educational materials [ASK GREG]
12.5 Other community resources on package-making:
You can find a diversity of additional helpful resources and tutorials on making R packages. We list a few below:
- Hilary Parker on basic package functions: bare-bones function making
- Writing an R package from scratch : Follows up on Hilary’s but incorporates usethis()
- Taking your data to Go: Straightforward, concise tutorial on data packages
- R Package Primer